QUESTION IMAGE
Question
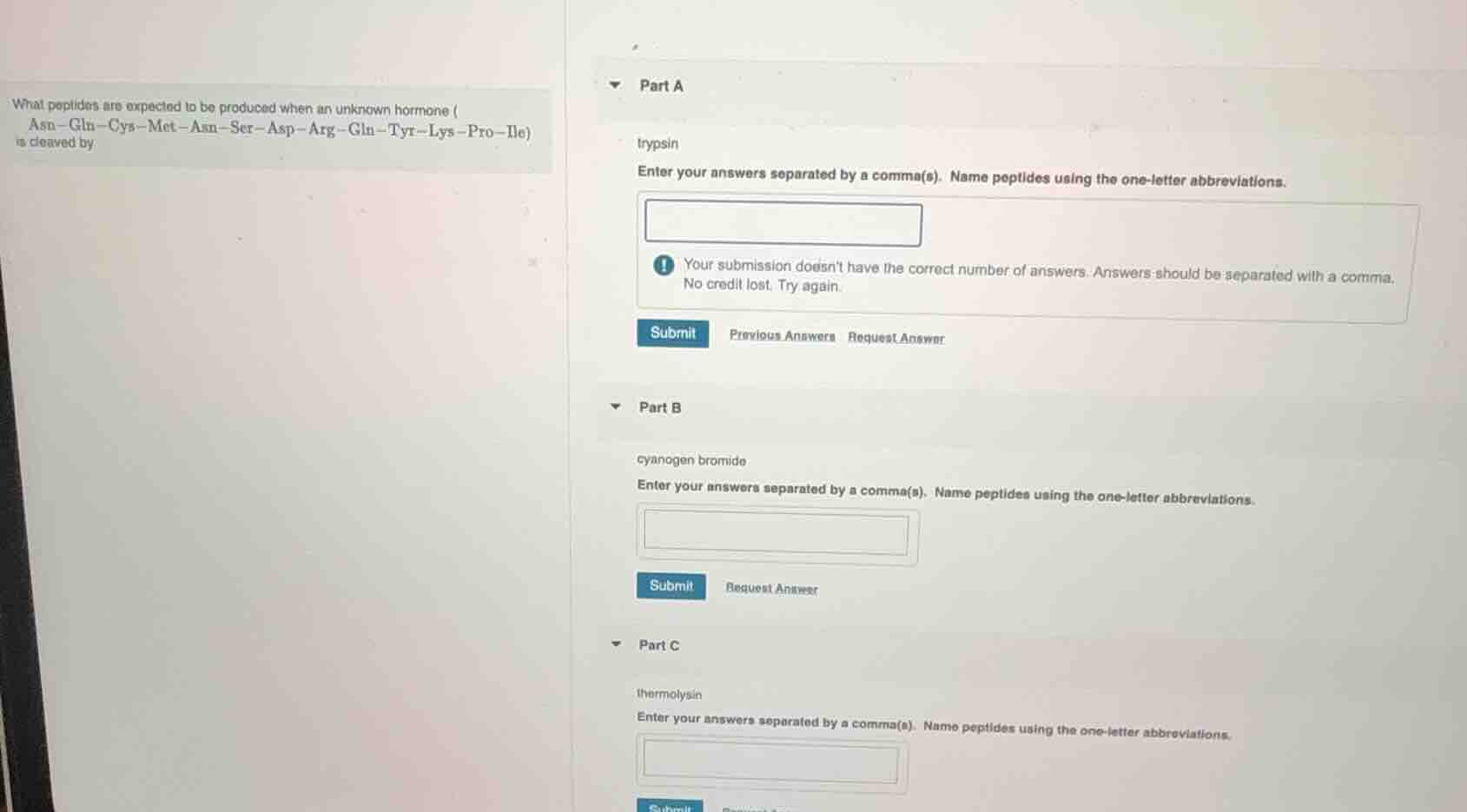
what peptides are expected to be produced when an unknown hormone (asn—gln—cys—met—asn—ser—asp—arg—gln—tyr—lys—pro—ile) is cleaved by
part a
trypsin
enter your answers separated by a comma(s). name peptides using the one - letter abbreviations.
your submission doesnt have the correct number of answers. answers should be separated with a comma.
no credit lost. try again.
submit previous answers request answer
part b
cyanogen bromide
enter your answers separated by a comma(s). name peptides using the one - letter abbreviations.
submit request answer
part c
thermolysin
enter your answers separated by a comma(s). name peptides using the one - letter abbreviations.
Part A (Trypsin Cleavage)
Step1: Recall Trypsin Cleavage Rule
Trypsin cleaves peptide bonds at the carboxyl side of Lys (K) and Arg (R), except when followed by Pro (P).
The sequence (one - letter: N - Q - C - M - N - S - D - R - Q - Y - K - P - I) has Arg (R) at position 8 and Lys (K) at position 11. But Lys (K) is followed by Pro (P), so trypsin won't cleave after K here.
- Cleave after R (position 8): First peptide: NQCMNSDR (positions 1 - 8), Second peptide: QYKPI (positions 9 - 13)? Wait, no, let's list the one - letter codes properly. The original sequence in one - letter: Asn (N), Gln (Q), Cys (C), Met (M), Asn (N), Ser (S), Asp (D), Arg (R), Gln (Q), Tyr (Y), Lys (K), Pro (P), Ile (I).
Trypsin cleaves at R (after R) and at K (but K is followed by P, so no cleavage at K). So after R (position 8, one - letter R), the first peptide is NQCMNSDR (residues 1 - 8: N, Q, C, M, N, S, D, R), and the second peptide is QYKPI (residues 9 - 13: Q, Y, K, P, I? Wait, no, residue 9 is Q, 10 is Y, 11 is K, 12 is P, 13 is I. But trypsin cleaves at the carboxyl of R and K (when not followed by P). Wait, maybe I made a mistake. Let's re - examine: The sequence is N - Q - C - M - N - S - D - R - Q - Y - K - P - I.
Trypsin cleaves peptide bonds on the C - terminal side of Lys (K) and Arg (R), unless the next residue is Pro (P). So:
- At R (position 8, one - letter R): the bond between R and Q (position 9) is cleaved? No, trypsin cleaves the bond after R (i.e., between R and the next amino acid). So the first peptide is NQCMNSDR (residues 1 - 8: N, Q, C, M, N, S, D, R), and the second peptide is QYKPI? Wait, no, residue 9 is Q, 10 is Y, 11 is K, 12 is P, 13 is I. But K is followed by P, so trypsin doesn't cleave after K. So the two peptides are NQCMNSDR, QYKPI? Wait, no, let's count the residues. The sequence has 13 residues? Wait, original sequence: Asn - Gln - Cys - Met - Asn - Ser - Asp - Arg - Gln - Tyr - Lys - Pro - Ile: that's 13 amino acids (1:Asn, 2:Gln, 3:Cys, 4:Met, 5:Asn, 6:Ser, 7:Asp, 8:Arg, 9:Gln, 10:Tyr, 11:Lys, 12:Pro, 13:Ile).
So after Arg (position 8), the first peptide is residues 1 - 8: N, Q, C, M, N, S, D, R (length 8). Then residues 9 - 13: Q, Y, K, P, I (length 5). But wait, is there a cleavage at K? K is at position 11, followed by P (position 12), so trypsin does not cleave after K. So the two peptides are NQCMNSDR, QYKPI? Wait, no, maybe I messed up the numbering. Let's write the one - letter sequence: N Q C M N S D R Q Y K P I.
Trypsin cleaves at R (after R) and at K (if not followed by P). So R is at position 8 (0 - based or 1 - based? Let's use 1 - based: position 8 is R. So cleave between position 8 (R) and 9 (Q): first peptide: 1 - 8 (N, Q, C, M, N, S, D, R), second peptide: 9 - 13 (Q, Y, K, P, I). But K is at position 11 (1 - based), followed by P (position 12), so no cleavage at K. So the two peptides are NQCMNSDR, QYKPI? Wait, but let's check again. Maybe the correct cleavage is:
Wait, the sequence is N (1), Q (2), C (3), M (4), N (5), S (6), D (7), R (8), Q (9), Y (10), K (11), P (12), I (13).
Trypsin cleaves at R (8) and at K (11) only if K is not followed by P. Since K (11) is followed by P (12), no cleavage at K. So only cleavage at R (8). So the two peptides are NQCMNSDR (1 - 8) and QYKPI (9 - 13). Wait, but maybe I made a mistake. Let's confirm the trypsin cleavage rule: Trypsin cleaves peptide bonds on the carboxyl side of lysine (Lys, K) or arginine (Arg, R), unless the next amino acid is proline (Pro, P). So in the sequence, after R (position 8, 1 - based), we cleave, and after K (position 11, 1 - based), since the next amino acid…
Cyanogen bromide (CNBr) cleaves at the carboxyl side of Met (M). The one - letter sequence is N Q C M N S D R Q Y K P I. Met (M) is at position 4 (1 - based). So cleave after M (position 4).
- First peptide: NQCM (residues 1 - 4: N, Q, C, M), Second peptide: NSDRQYKPI (residues 5 - 13: N, S, D, R, Q, Y, K, P, I).
Thermolysin cleaves at the amino side of hydrophobic amino acids (Leu, Ile, Val, Met, Phe, Trp, Tyr). The one - letter sequence: N (polar), Q (polar), C (polar), M (hydrophobic), N (polar), S (polar), D (polar), R (polar), Q (polar), Y (hydrophobic), K (polar), P (polar), I (hydrophobic).
Hydrophobic residues: M (4), Y (10), I (13).
- Cleave before M (position 4): First peptide: NQC (residues 1 - 3: N, Q, C), Second peptide: MNSDRQYKPI (residues 4 - 13: M, N, S, D, R, Q, Y, K, P, I)? No, thermolysin cleaves at the amino side of hydrophobic residues, i.e., the bond before the hydrophobic residue. So for M (position 4, hydrophobic), cleave before M: peptide 1: NQC (1 - 3), peptide 2: MNSDRQYKPI (4 - 13). For Y (position 10, hydrophobic), cleave before Y: peptide 2 is split into MNSDRQ (4 - 9) and YKPI (10 - 13). For I (position 13, hydrophobic), cleave before I: but I is the last residue, so no cleavage before I.
Wait, let's do it step by step. The hydrophobic residues are M (4), Y (10), I (13).
- Cleave before M (4): peptide 1: NQC (1 - 3), peptide 2: MNSDRQYKPI (4 - 13).
- Then cleave before Y (10) in peptide 2: peptide 2a: MNSDRQ (4 - 9), peptide 2b: YKPI (10 - 13).
- Then cleave before I (13) in peptide 2b: but I is the last residue, so no cleavage.
So the peptides are NQC, MNSDRQ, YKPI. Wait, let's check the hydrophobic residues again. Thermolysin cleaves at the amino side of hydrophobic amino acids (so the bond between the residue before the hydrophobic residue and the hydrophobic residue is cleaved). So:
- Before M (M is at 4, one - letter M), the residue before M is C (position 3, C is polar? Wait, no, thermolysin cleaves at the N - terminal side of hydrophobic amino acids (i.e., the bond between the (n - 1)th and nth residue, where nth is hydrophobic). So for M (n = 4), the (n - 1)th residue is C (position 3, C is polar), so cleave between C (3) and M (4): peptide 1: NQC (1 - 3), peptide 2: MNSDRQYKPI (4 - 13).
For Y (n = 10, Y is hydrophobic), the (n - 1)th residue is Q (position 9, Q is polar), so cleave between Q (9) and Y (10): peptide 2a: MNSDRQ (4 - 9), peptide 2b: YKPI (10 - 13).
For I (n = 13, I is hydrophobic), the (n - 1)th residue is P (position 12, P is polar), so cleave between P (12) and I (13): peptide 2b is split into YKP (10 - 12) and I (13)? Wait, no, thermolysin cleaves at the amino side of hydrophobic residues, so the bond before the hydrophobic residue. So for I (position 13), the bond before I is between P (12) and I (13), so cleave there. So now:
- After cleaving before M: NQC, MNSDRQYKPI
- After cleaving before Y: NQC, MNSDRQ, YKPI
- After cleaving before I: NQC, MNSDRQ, YKP, I
Wait, but let's check the hydrophobic residues: M (4), Y (10), I (13). So three hydrophobic residues, so three cleavages (before each hydrophobic residue, except if it's the first residue). So:
- Before M (4): cleave between C (3) and M (4): peptide 1: NQC (1 - 3)
- Before Y (10): cleave between Q (9) and Y (10): peptide 2: MNSDRQ (4 - 9), peptide 3: YKPI (10 - 13)
- Before I (13): cleave between P (12) and I (13): peptide 3 is split into YKP (10 - 12) and I (13)
Wait, but maybe a simpler way: thermolysin cleaves at the N - terminal side of hydrophobic amino acids, so the peptides are formed by cleaving before M, Y, and I.
So the peptides are:
- NQC (before M: residues 1 - 3)
- MNSDRQ (before Y: residues 4 - 9)
- YKP (before I: residues 10 - 12)
- I (residue 13)
But let's check the sequence length. Original sequence has 13 residues. NQC (3) + MNSDRQ (6) + YKP (3) + I (1) = 3 + 6+3 + 1 = 13. Yes.
Snap & solve any problem in the app
Get step-by-step solutions on Sovi AI
Photo-based solutions with guided steps
Explore more problems and detailed explanations
(Part A): NQCMNSDR, QYKPI