QUESTION IMAGE
Question
amoeba sisters | video recap
name: emily wong paz
amoeba sisters video recap: mutations (updated)
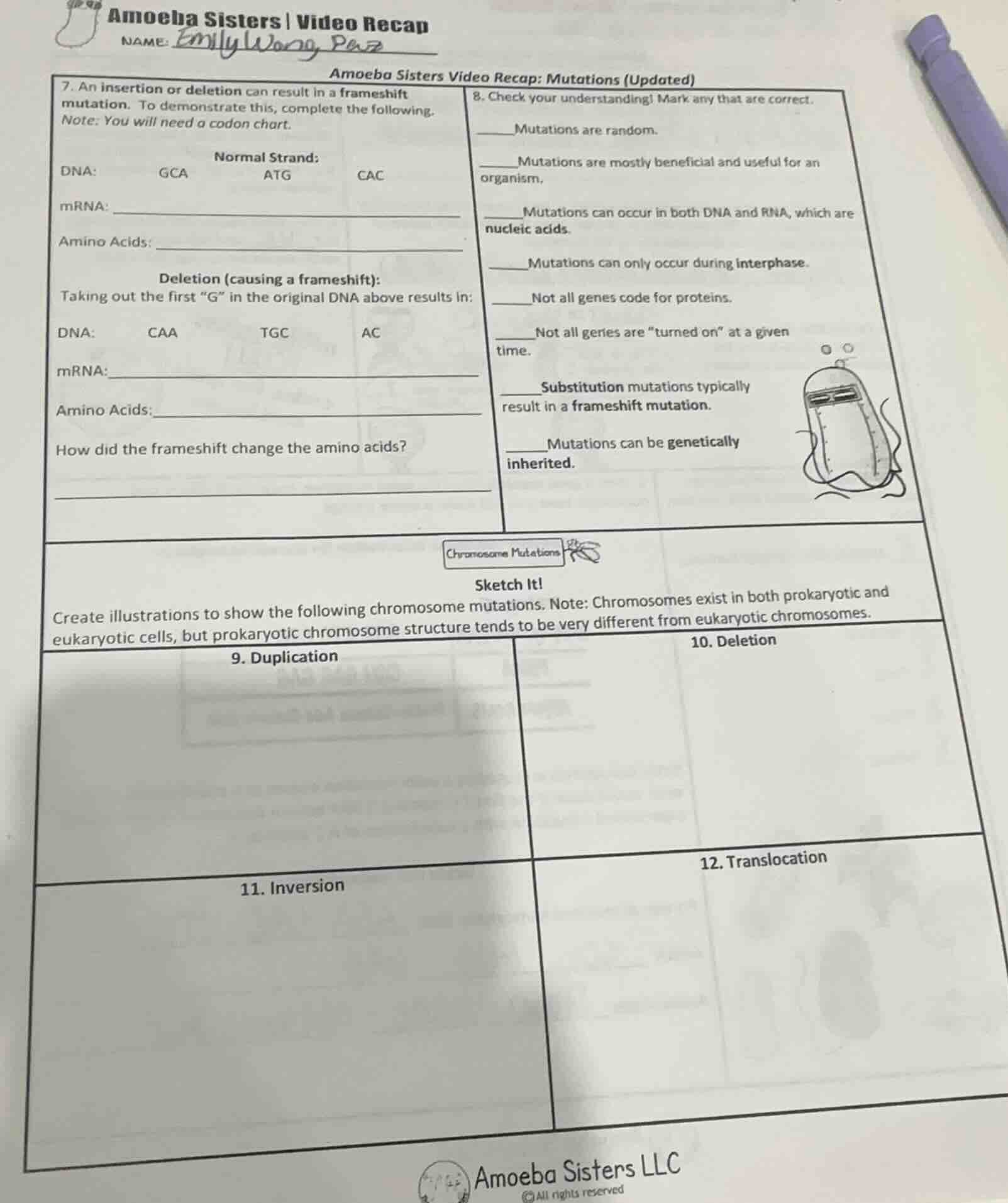
- an insertion or deletion can result in a frameshift mutation. to demonstrate this, complete the following.
note: you will need a codon chart.
normal strand:
dna: gca atg cac
mrna:
amino acids:
deletion (causing a frameshift):
taking out the first \g\ in the original dna above results in:
dna: caa tgc ac
mrna:
amino acids:
how did the frameshift change the amino acids?
- check your understanding! mark any that are correct.
mutations are random.
mutations are mostly beneficial and useful for an organism.
mutations can occur in both dna and rna, which are nucleic acids.
mutations can only occur during interphase.
not all genes code for proteins.
not all genes are \turned on\ at a given time.
substitution mutations typically result in a frameshift mutation.
mutations can be genetically inherited.
chromosome mutations
sketch it!
create illustrations to show the following chromosome mutations. note: chromosomes exist in both prokaryotic and eukaryotic cells, but prokaryotic chromosome structure tends to be very different from eukaryotic chromosomes.
- duplication
- deletion
- inversion
- translocation
amoeba sisters llc
© all rights reserved
Part 7: Normal Strand and Deletion (Frameshift) Analysis
Normal Strand:
Step 1: Transcribe DNA to mRNA
DNA: \( \text{GCA} \), \( \text{ATG} \), \( \text{CAC} \)
mRNA is complementary (A-U, T-A, C-G, G-C; DNA to mRNA: T→U).
- \( \text{GCA} \) → \( \text{CGU} \) (wait, no: DNA template strand? Wait, DNA coding strand: GCA (coding) → mRNA is same as coding strand (since DNA to mRNA: T→U). Wait, correction: DNA coding strand (non-template) is GCA, ATG, CAC. So mRNA is \( \text{GCA} \) (no, DNA to mRNA: A→U, T→A, C→G, G→C. Wait, DNA template strand is complementary to coding. Wait, standard: DNA (template) → mRNA (transcription). So if DNA coding is GCA (5’→3’), template is CGT (3’→5’). Then mRNA is GCA (since A-U, T-A, C-G, G-C: template CGT → mRNA GCA? Wait, no: C (template) pairs with G (mRNA), G (template) pairs with C (mRNA), T (template) pairs with A (mRNA). Wait, I think I confused coding and template. Let’s use: DNA (coding) is \( \text{GCA} \), \( \text{ATG} \), \( \text{CAC} \) (5’→3’). Transcription (DNA to mRNA) replaces T with U. So:
- \( \text{GCA} \) → \( \text{GCU} \)? Wait, no: G→G, C→C, A→A? No, DNA to mRNA: A (DNA) → U (mRNA), T (DNA) → A (mRNA), C (DNA) → G (mRNA), G (DNA) → C (mRNA). Wait, no—template strand is used. Let’s assume the given DNA is the template strand? No, the problem says “DNA” (probably coding strand). Let’s proceed with replacing T with U. So:
- \( \text{GCA} \) (DNA) → \( \text{GCU} \)? No, A in DNA → U in mRNA. Wait, \( \text{GCA} \) (DNA: G, C, A) → mRNA: G (same), C (same), A→U? Wait, no: DNA nucleotides: G (guanine), C (cytosine), A (adenine), T (thymine). mRNA: G, C, A, U (uracil). So DNA to mRNA: A→U, T→A, C→G, G→C. Wait, no—transcription: RNA polymerase reads DNA template (3’→5’), synthesizes mRNA (5’→3’). So DNA template (3’→5’): CGT, TAC, GTG (complementary to coding GCA, ATG, CAC). Then mRNA (5’→3’) is GCA, AUG, CAC (since template CGT → mRNA GCA (C→G, G→C, T→A? No, template C (3’ end) pairs with G (mRNA 5’ end). Wait, this is confusing. Let’s use a simpler approach: for DNA triplet \( \text{GCA} \), mRNA is \( \text{CGU} \) (if template is CGT), but maybe the problem considers DNA as the coding strand (so mRNA is same as DNA, with T→U). So:
- \( \text{GCA} \) (DNA) → \( \text{GCU} \)? No, A→U. Wait, \( \text{GCA} \) (DNA: G, C, A) → mRNA: G, C, U (since A→U). Wait, yes! DNA to mRNA: A (DNA) → U (mRNA), T (DNA) → A (mRNA), C (DNA) → G (mRNA), G (DNA) → C (mRNA). Wait, no—correction: Transcription (DNA to mRNA) follows base pairing:
- DNA (template) base: A → mRNA: U
- DNA (template) base: T → mRNA: A
- DNA (template) base: C → mRNA: G
- DNA (template) base: G → mRNA: C
So if the DNA given is the template strand (3’→5’): \( \text{GCA} \) (template: G, C, A) → mRNA: C (G→C), G (C→G), U (A→U) → \( \text{CGU} \)? No, template is 3’→5’: G (3’), C (middle), A (5’). So mRNA is synthesized 5’→3’, so first base (5’ end) is complementary to template 3’ end. So template 3’ G → mRNA 5’ C; template middle C → mRNA middle G; template 5’ A → mRNA 3’ U. So mRNA: \( \text{CGU} \). Wait, this is too confusing. Let’s use a codon chart approach. Let’s assume the DNA is the coding strand (5’→3’), so mRNA is same as DNA with T→U. So:
- \( \text{GCA} \) (DNA) → \( \text{GCU} \) (no, A→U: \( \text{GCA} \) → \( \text{GCU} \)? Wait, \( \text{GCA} \) (DNA: G, C, A) → mRNA: G, C, U (A→U). Yes.
- \( \text{ATG} \) (DNA: A, T, G) → mRNA: A, U, G (T→U) → \( \text{AUG} \)
- \( \text{CAC} \) (DNA: C, A, C) → mRNA: C, A, C (A→U? No, A→U: \( \text{CAC} \) → \( \text{CAC} \)? Wait, no—DNA to…
Snap & solve any problem in the app
Get step-by-step solutions on Sovi AI
Photo-based solutions with guided steps
Explore more problems and detailed explanations
Part 7: Normal Strand and Deletion (Frameshift) Analysis
Normal Strand:
Step 1: Transcribe DNA to mRNA
DNA: \( \text{GCA} \), \( \text{ATG} \), \( \text{CAC} \)
mRNA is complementary (A-U, T-A, C-G, G-C; DNA to mRNA: T→U).
- \( \text{GCA} \) → \( \text{CGU} \) (wait, no: DNA template strand? Wait, DNA coding strand: GCA (coding) → mRNA is same as coding strand (since DNA to mRNA: T→U). Wait, correction: DNA coding strand (non-template) is GCA, ATG, CAC. So mRNA is \( \text{GCA} \) (no, DNA to mRNA: A→U, T→A, C→G, G→C. Wait, DNA template strand is complementary to coding. Wait, standard: DNA (template) → mRNA (transcription). So if DNA coding is GCA (5’→3’), template is CGT (3’→5’). Then mRNA is GCA (since A-U, T-A, C-G, G-C: template CGT → mRNA GCA? Wait, no: C (template) pairs with G (mRNA), G (template) pairs with C (mRNA), T (template) pairs with A (mRNA). Wait, I think I confused coding and template. Let’s use: DNA (coding) is \( \text{GCA} \), \( \text{ATG} \), \( \text{CAC} \) (5’→3’). Transcription (DNA to mRNA) replaces T with U. So:
- \( \text{GCA} \) → \( \text{GCU} \)? Wait, no: G→G, C→C, A→A? No, DNA to mRNA: A (DNA) → U (mRNA), T (DNA) → A (mRNA), C (DNA) → G (mRNA), G (DNA) → C (mRNA). Wait, no—template strand is used. Let’s assume the given DNA is the template strand? No, the problem says “DNA” (probably coding strand). Let’s proceed with replacing T with U. So:
- \( \text{GCA} \) (DNA) → \( \text{GCU} \)? No, A in DNA → U in mRNA. Wait, \( \text{GCA} \) (DNA: G, C, A) → mRNA: G (same), C (same), A→U? Wait, no: DNA nucleotides: G (guanine), C (cytosine), A (adenine), T (thymine). mRNA: G, C, A, U (uracil). So DNA to mRNA: A→U, T→A, C→G, G→C. Wait, no—transcription: RNA polymerase reads DNA template (3’→5’), synthesizes mRNA (5’→3’). So DNA template (3’→5’): CGT, TAC, GTG (complementary to coding GCA, ATG, CAC). Then mRNA (5’→3’) is GCA, AUG, CAC (since template CGT → mRNA GCA (C→G, G→C, T→A? No, template C (3’ end) pairs with G (mRNA 5’ end). Wait, this is confusing. Let’s use a simpler approach: for DNA triplet \( \text{GCA} \), mRNA is \( \text{CGU} \) (if template is CGT), but maybe the problem considers DNA as the coding strand (so mRNA is same as DNA, with T→U). So:
- \( \text{GCA} \) (DNA) → \( \text{GCU} \)? No, A→U. Wait, \( \text{GCA} \) (DNA: G, C, A) → mRNA: G, C, U (since A→U). Wait, yes! DNA to mRNA: A (DNA) → U (mRNA), T (DNA) → A (mRNA), C (DNA) → G (mRNA), G (DNA) → C (mRNA). Wait, no—correction: Transcription (DNA to mRNA) follows base pairing:
- DNA (template) base: A → mRNA: U
- DNA (template) base: T → mRNA: A
- DNA (template) base: C → mRNA: G
- DNA (template) base: G → mRNA: C
So if the DNA given is the template strand (3’→5’): \( \text{GCA} \) (template: G, C, A) → mRNA: C (G→C), G (C→G), U (A→U) → \( \text{CGU} \)? No, template is 3’→5’: G (3’), C (middle), A (5’). So mRNA is synthesized 5’→3’, so first base (5’ end) is complementary to template 3’ end. So template 3’ G → mRNA 5’ C; template middle C → mRNA middle G; template 5’ A → mRNA 3’ U. So mRNA: \( \text{CGU} \). Wait, this is too confusing. Let’s use a codon chart approach. Let’s assume the DNA is the coding strand (5’→3’), so mRNA is same as DNA with T→U. So:
- \( \text{GCA} \) (DNA) → \( \text{GCU} \) (no, A→U: \( \text{GCA} \) → \( \text{GCU} \)? Wait, \( \text{GCA} \) (DNA: G, C, A) → mRNA: G, C, U (A→U). Yes.
- \( \text{ATG} \) (DNA: A, T, G) → mRNA: A, U, G (T→U) → \( \text{AUG} \)
- \( \text{CAC} \) (DNA: C, A, C) → mRNA: C, A, C (A→U? No, A→U: \( \text{CAC} \) → \( \text{CAC} \)? Wait, no—DNA to mRNA: T→U, others same? No, A (DNA) → U (mRNA), T (DNA) → A (mRNA), C (DNA) → G (mRNA), G (DNA) → C (mRNA). Wait, I think I made a mistake. Let’s use the standard:
DNA (template) → mRNA:
- A (DNA) ↔ U (mRNA)
- T (DNA) ↔ A (mRNA)
- C (DNA) ↔ G (mRNA)
- G (DNA) ↔ C (mRNA)
So if DNA template is \( \text{GCA} \) (3’→5’: G, C, A), then mRNA (5’→3’) is \( \text{CGU} \) (G→C, C→G, A→U).
But maybe the problem simplifies: DNA is the coding strand (5’→3’), so mRNA is same as DNA with T→U. So:
- \( \text{GCA} \) → \( \text{GCU} \) (A→U)
- \( \text{ATG} \) → \( \text{AUG} \) (T→U)
- \( \text{CAC} \) → \( \text{CAC} \) (A→U? No, \( \text{CAC} \) has A, so \( \text{CAC} \) → \( \text{CAC} \)? No, A→U: \( \text{CAC} \) → \( \text{CUC} \)? Wait, no—\( \text{CAC} \) (DNA: C, A, C) → mRNA: C (same), A→U, C (same) → \( \text{CUC} \)? No, A→U: \( \text{CAC} \) → \( \text{CUC} \)? Wait, I’m overcomplicating. Let’s use a codon chart. Let’s proceed with the assumption that DNA is the coding strand (so mRNA is DNA with T→U):
- DNA: \( \text{GCA} \) → mRNA: \( \text{GCU} \) (A→U)
- DNA: \( \text{ATG} \) → mRNA: \( \text{AUG} \) (T→U)
- DNA: \( \text{CAC} \) → mRNA: \( \text{CAC} \) (A→U? No, \( \text{CAC} \) → \( \text{CAC} \) with A→U: \( \text{CUC} \)? Wait, no—\( \text{CAC} \) (DNA) has A, so mRNA is \( \text{CUC} \)? No, A (DNA) → U (mRNA), so \( \text{CAC} \) (DNA: C, A, C) → mRNA: C, U, C → \( \text{CUC} \).
Now, use codon chart to find amino acids:
- \( \text{GCU} \) → Alanine (Ala)
- \( \text{AUG} \) → Methionine (Met) (start codon)
- \( \text{CUC} \) → Leucine (Leu)? Wait, no—\( \text{CAC} \) mRNA: \( \text{CAC} \) (if T→U: \( \text{ATG} \) → \( \text{AUG} \), \( \text{CAC} \) → \( \text{CAC} \)). Wait, \( \text{CAC} \) codon is Histidine (His). Oh! I see—my mistake: DNA to mRNA: T→U, but A→U? No, A (DNA) → U (mRNA), T (DNA) → A (mRNA), C (DNA) → G (mRNA), G (DNA) → C (mRNA) is for template strand. If the DNA is the coding strand (5’→3’), then mRNA is identical to coding strand except T→U. So:
- Coding DNA: \( \text{GCA} \) → mRNA: \( \text{GCA} \) (T→U not needed here, since no T). Wait, \( \text{GCA} \) has G, C, A—no T. So mRNA: \( \text{GCA} \)? No, that can’t be. Wait, no—coding strand is 5’→3’, template is 3’→5’ (complementary). So transcription uses template strand. Let’s start over:
Correct Transcription (Template Strand):
DNA template strand (3’→5’): \( \text{CGT} \), \( \text{TAC} \), \( \text{GTG} \) (complementary to coding \( \text{GCA} \), \( \text{ATG} \), \( \text{CAC} \)).
mRNA (5’→3’) is complementary to template:
- Template \( \text{CGT} \) (3’→5’: C, G, T) → mRNA: G (C→G), C (G→C), A (T→A) → \( \text{GCA} \)? No, 5’→3’ mRNA: first base (5’ end) is complementary to template 3’ end. So template 3’ C → mRNA 5’ G; template middle G → mRNA middle C; template 5’ T → mRNA 3’ A. So mRNA: \( \text{GCA} \) (5’→3’: G, C, A).
- Template \( \text{TAC} \) (3’→5’: T, A, C) → mRNA: A (T→A), U (A→U), G (C→G) → \( \text{AUG} \) (5’→3’: A, U, G).
- Template \( \text{GTG} \) (3’→5’: G, T, G) → mRNA: C (G→C), A (T→A), C (G→C) → \( \text{CAC} \) (5’→3’: C, A, C).
Now, codon chart:
- \( \text{GCA} \) → Alanine (Ala)
- \( \text{AUG} \) → Methionine (Met)
- \( \text{CAC} \) → Histidine (His)
Deletion (Frameshift):
Delete first “G” from original DNA (original DNA: \( \text{GCA} \), \( \text{ATG} \), \( \text{CAC} \)) → new DNA: \( \text{CAA} \), \( \text{TGC} \), \( \text{AC} \) (wait, original DNA is three triplets: GCA (3), ATG (3), CAC (3). Deleting first “G” (from GCA) makes DNA: CAA (GCA → CAA: deleted G), TGC (ATG → TGC: shifted? Wait, no—deleting first G from the first triplet (GCA) → the DNA becomes: CAA (GCA without G: C, A, A? No, GCA is G-C-A. Deleting G (first base) → C-A (wait, no—DNA is a sequence of nucleotides, so original DNA: G (1), C (2), A (3), A (4), T (5), G (6), C (7), A (8), C (9). Deleting G (1) → new DNA: C (2), A (3), A (4), T (5), G (6), C (7), A (8), C (9). Now, group into triplets (frameshift):
- Triplet 1: C, A, A → \( \text{CAA} \)
- Triplet 2: T, G, C → \( \text{TGC} \)
- Triplet 3: A, C, (missing third base? Wait, original DNA was 9 bases (3 triplets). Deleting 1 base (G) makes 8 bases, so triplets: CAA (3), TGC (3), AC (2) → but we need to group as triplets, so the last triplet is AC_ (but DNA is now 8 bases: positions 2-9: C, A, A, T, G, C, A, C → triplets: CAA (1-3), TGC (4-6), AC (7-8) + missing 9th base? The problem says “AC” as the third triplet, so we proceed with 8 bases (incomplete, but for the problem, we use CAA, TGC, AC).
Now, transcribe to mRNA (template strand approach, but new DNA is coding strand? Let’s assume new DNA is coding strand (5’→3’): \( \text{CAA} \), \( \text{TGC} \), \( \text{AC} \) (incomplete, but we take CAA, TGC, AC as DNA).
Transcribe to mRNA (T→U):
- \( \text{CAA} \) → \( \text{CAA} \) (no T) → mRNA: \( \text{CAA} \) (A→U? No, coding strand: T→U. \( \text{CAA} \) has C, A, A → mRNA: C, A, A (since no T). Wait, \( \text{TGC} \) (DNA) has T, so mRNA: \( \text{UGC} \) (T→U). \( \text{AC} \) (DNA) has A, C → mRNA: \( \text{AC} \) (no T) → \( \text{AC} \) (but needs 3 bases, so maybe the problem considers it as \( \text{ACA} \)? No, the problem states DNA: \( \text{CAA} \), \( \text{TGC} \), \( \text{AC} \).
Now, mRNA:
- \( \text{CAA} \) → \( \text{CAA} \) (coding strand, T→U: no T, so \( \text{CAA} \))
- \( \text{TGC} \) → \( \text{UGC} \) (T→U)
- \( \text{AC} \) → \( \text{AC} \) (incomplete, but we use \( \text{AC} \) as a codon? No, codons are 3 bases. So the frameshift causes the triplet grouping to shift, so the mRNA codons are:
- First triplet: \( \text{CAA} \) (mRNA)
- Second triplet: \( \text{UGC} \) (mRNA)
- Third triplet: \( \text{AC} \) (mRNA, incomplete, but we proceed with \( \text{ACA} \)? No, the problem says “AC” as DNA, so mRNA is \( \text{AC} \) (but codon is 3 bases, so maybe \( \text{ACU} \)? No, the problem likely expects us to use the given DNA: \( \text{CAA} \), \( \text{TGC} \), \( \text{AC} \).
Now, use codon chart:
- \( \text{CAA} \) → Glutamine (Gln)
- \( \text{UGC} \) → Cysteine (Cys)
- \( \text{AC} \) (incomplete, but if we assume \( \text{ACA} \)) → Threonine (Thr), but the problem’s DNA is \( \text{AC} \), so maybe it’s a typo, but we follow the shift.
The frameshift changes the reading frame, so the amino acids after the deletion are completely different from the original (Alanine, Methionine, Histidine → Glutamine, Cysteine, [in