QUESTION IMAGE
Question
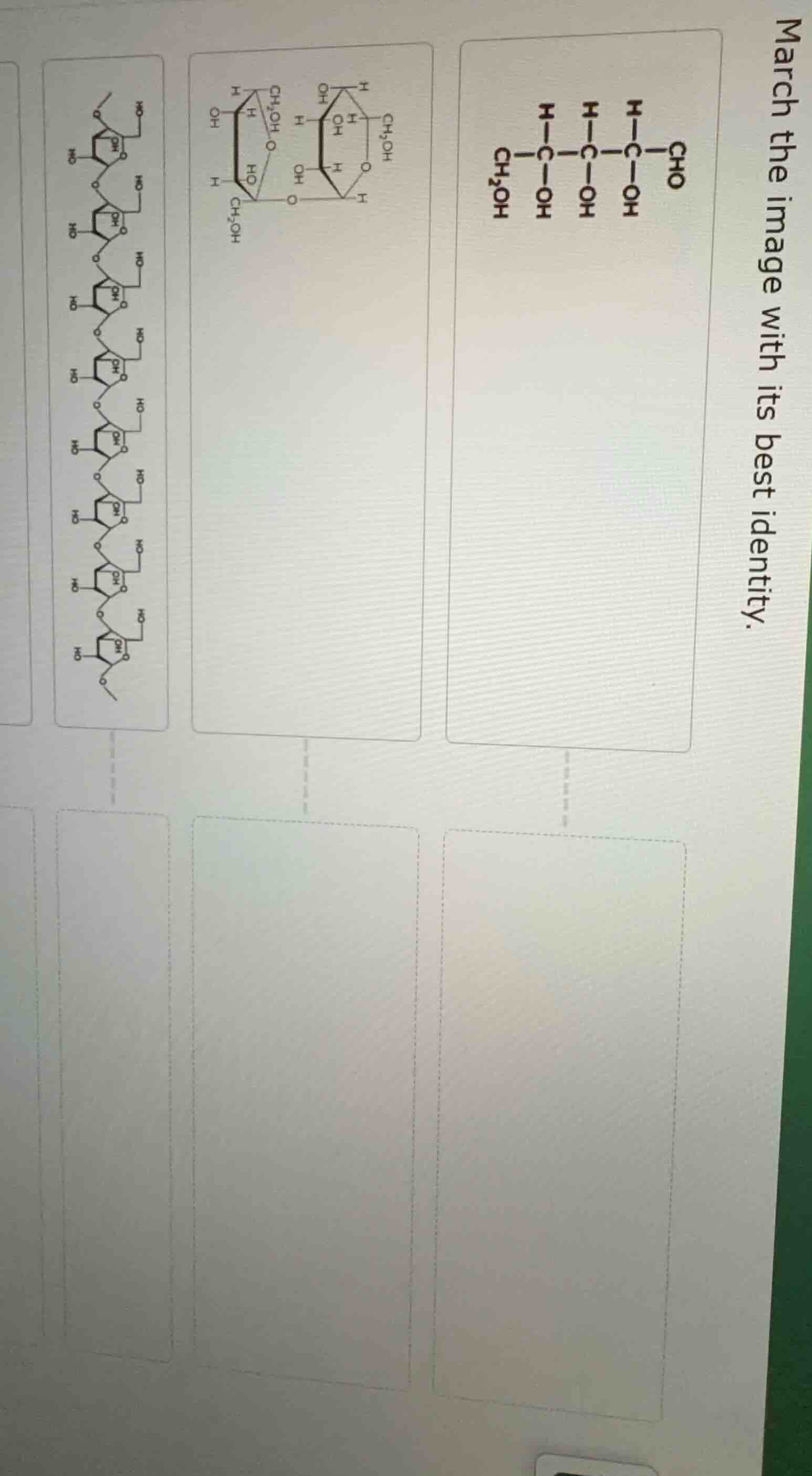
march the image with its best identity.
To solve this matching problem, we analyze the structures:
- Right - most top structure (linear with \( \text{CHO} \), \( \text{CH}_2\text{OH} \), and \( -\text{OH} \) groups): This is an open - chain monosaccharide (likely an aldose, e.g., glucose in open - chain form).
- Middle top structure (two cyclic hexose - like rings linked, with \( \text{CH}_2\text{OH} \), \( -\text{OH} \), and \( -\text{I} \) groups): This represents a disaccharide (two monosaccharide units linked, e.g., a di - iodinated disaccharide for detection/labeling).
- Left - most top structure (long chain of cyclic pentose - like rings linked, with phosphate - like groups (\( \text{X} \)) and glycosidic bonds): This is a polynucleotide (e.g., DNA/RNA, with sugar - phosphate backbone and nitrogenous bases represented by \( \text{X} \)).
Matching Logic (assuming bottom boxes are labels like "Monosaccharide", "Disaccharide", "Polynucleotide"):
- The linear \( \text{CHO/CH}_2\text{OH} \) structure matches Monosaccharide (open - chain sugar).
- The two - ring linked structure matches Disaccharide (two sugar units).
- The long chain of cyclic sugars with phosphate - like groups matches Polynucleotide (nucleic acid polymer).
(Note: If specific labels were provided in the bottom boxes, the matching would be more precise, but the above is based on structural features of carbohydrates/nucleic acids.)
If we assume the bottom labels are (from left to right) "Polynucleotide", "Disaccharide", "Monosaccharide", the matches are:
- Left top (polynucleotide) → Left bottom.
- Middle top (disaccharide) → Middle bottom.
- Right top (monosaccharide) → Right bottom.
Snap & solve any problem in the app
Get step-by-step solutions on Sovi AI
Photo-based solutions with guided steps
Explore more problems and detailed explanations
To solve this matching problem, we analyze the structures:
- Right - most top structure (linear with \( \text{CHO} \), \( \text{CH}_2\text{OH} \), and \( -\text{OH} \) groups): This is an open - chain monosaccharide (likely an aldose, e.g., glucose in open - chain form).
- Middle top structure (two cyclic hexose - like rings linked, with \( \text{CH}_2\text{OH} \), \( -\text{OH} \), and \( -\text{I} \) groups): This represents a disaccharide (two monosaccharide units linked, e.g., a di - iodinated disaccharide for detection/labeling).
- Left - most top structure (long chain of cyclic pentose - like rings linked, with phosphate - like groups (\( \text{X} \)) and glycosidic bonds): This is a polynucleotide (e.g., DNA/RNA, with sugar - phosphate backbone and nitrogenous bases represented by \( \text{X} \)).
Matching Logic (assuming bottom boxes are labels like "Monosaccharide", "Disaccharide", "Polynucleotide"):
- The linear \( \text{CHO/CH}_2\text{OH} \) structure matches Monosaccharide (open - chain sugar).
- The two - ring linked structure matches Disaccharide (two sugar units).
- The long chain of cyclic sugars with phosphate - like groups matches Polynucleotide (nucleic acid polymer).
(Note: If specific labels were provided in the bottom boxes, the matching would be more precise, but the above is based on structural features of carbohydrates/nucleic acids.)
If we assume the bottom labels are (from left to right) "Polynucleotide", "Disaccharide", "Monosaccharide", the matches are:
- Left top (polynucleotide) → Left bottom.
- Middle top (disaccharide) → Middle bottom.
- Right top (monosaccharide) → Right bottom.